Models of cell membranes reveal workings of biology’s greatest gatekeeper
 Mikko Haataja, assistant professor of mechanical and aerospace engineering, views cell membranes as “very complicated beasts.” As the beasts can’t always be explored experimentally, he uses computer simulations to probe deep into their structure and function.
Mikko Haataja, assistant professor of mechanical and aerospace engineering, views cell membranes as “very complicated beasts.” As the beasts can’t always be explored experimentally, he uses computer simulations to probe deep into their structure and function.
A heightened understanding of the structures-which control cell-to-cell communication and act as gatekeepers, determining what enters and exits the cell-will add to existing knowledge of disease processes and guide the development of more effective pharmaceutical drugs.
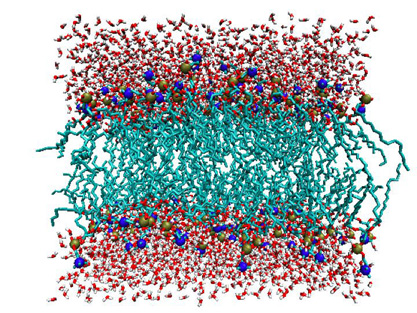
Haataja works with Athanassios Panagiotopoulos, the Susan Dod Brown Professor of Chemical Engineering, to model the architecture of cell membranes, which are made of lipids (i.e., fat or cholesterol) and protein. By poking and prodding these virtual models, Haataja explores how different factors affect the flow of substances and information into and out of the cell.
“In biology, there are multiple parameters we don’t understand, knobs we don’t know how to tune,” Haataja explained. “Computational materials science allows us to construct models and turn these knobs at will.”
He is currently investigating factors that might regulate the formation of lipid clusters that are hypothesized to exist in membranes. For instance, he predicted how the stability of these “lipid rafts” would be affected if molecules from inside the cell were able to join and leave the clusters. In future work, Haataja will collaborate with biologists to conduct wet lab experiments based on his findings.
As the rafts are thought to play an important role in the body’s immune response, a heightened understanding of their behavior could lead to better treatments for a number of diseases, including HIV and malaria.







