Trying to unravel the roles that a small set of genes play in the regulation of a human trait is a daunting enough task, but when scientists try to apply the same analytic methods to a specific tissue or organ, they quickly run into a storm of information.
The functional role of any one gene is quickly obscured by a cascade of genes whose influence combines with that of other genes and environmental factors to affect multiple pathways. What starts as a few bits of information quickly becomes a blizzard of complex structured interactions.
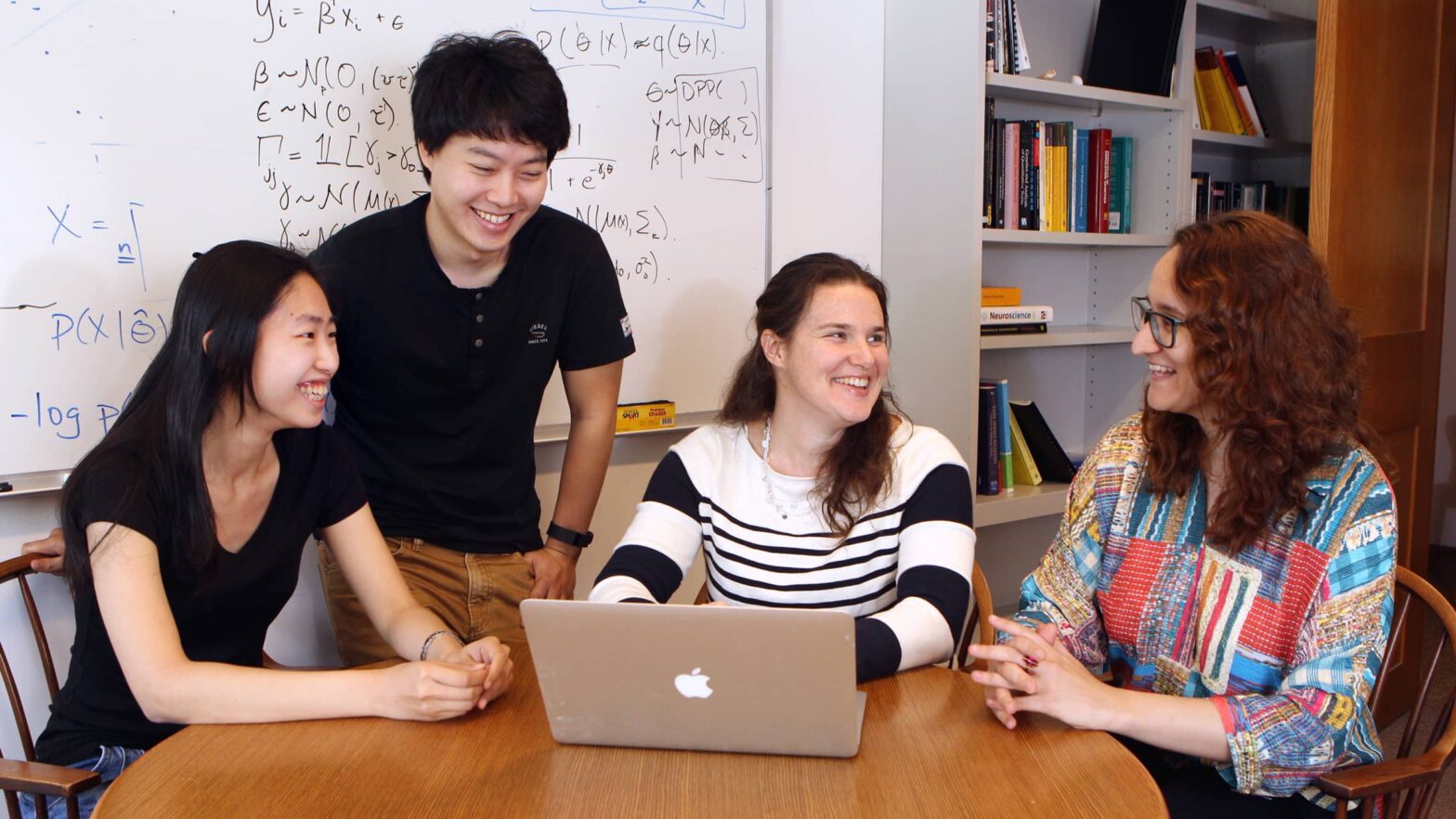
“It’s really a needle in a haystack,” said Barbara Engelhardt, an assistant professor of computer science.
Engelhardt’s research group specializes in handling statistically difficult problems such as deciphering gene expression data, as well as data about other complex traits and disease. One of the great challenges of her work is finding subtle patterns among relatively small numbers of samples – even a large clinical trial might involve only a few thousand people, but tens of thousands of genes and millions of genetic mutations.
“Statistics in general has been developed for very large data sets – a billion Facebook users, a trillion Google queries,” Engelhardt said. In most biological systems, on the other hand, “we don’t have an infinite amount of samples because of the cost to acquire each one.”
With relatively small pools of data, it can be difficult to separate biologically important patterns from random noise or technical effects. To address these problems, Engelhardt’s team harnesses the power of computation with innovative statistical methods. Identifying and characterizing these latent patterns can help scientists understand key mechanistic relationships in immense fields of data. Recent work has looked at the interactions of multiple genes in breast cancer, neuropsychiatric diseases and metabolic diseases.
“We are developing statistical methods to separate the structured noise from the signal,” she said. “We want to figure out the interactions driving disease from subtle patterns within the data.”