src=”/wp-content/uploads/treating-main-01.jpg” style=”width: 685px; height: 300px;
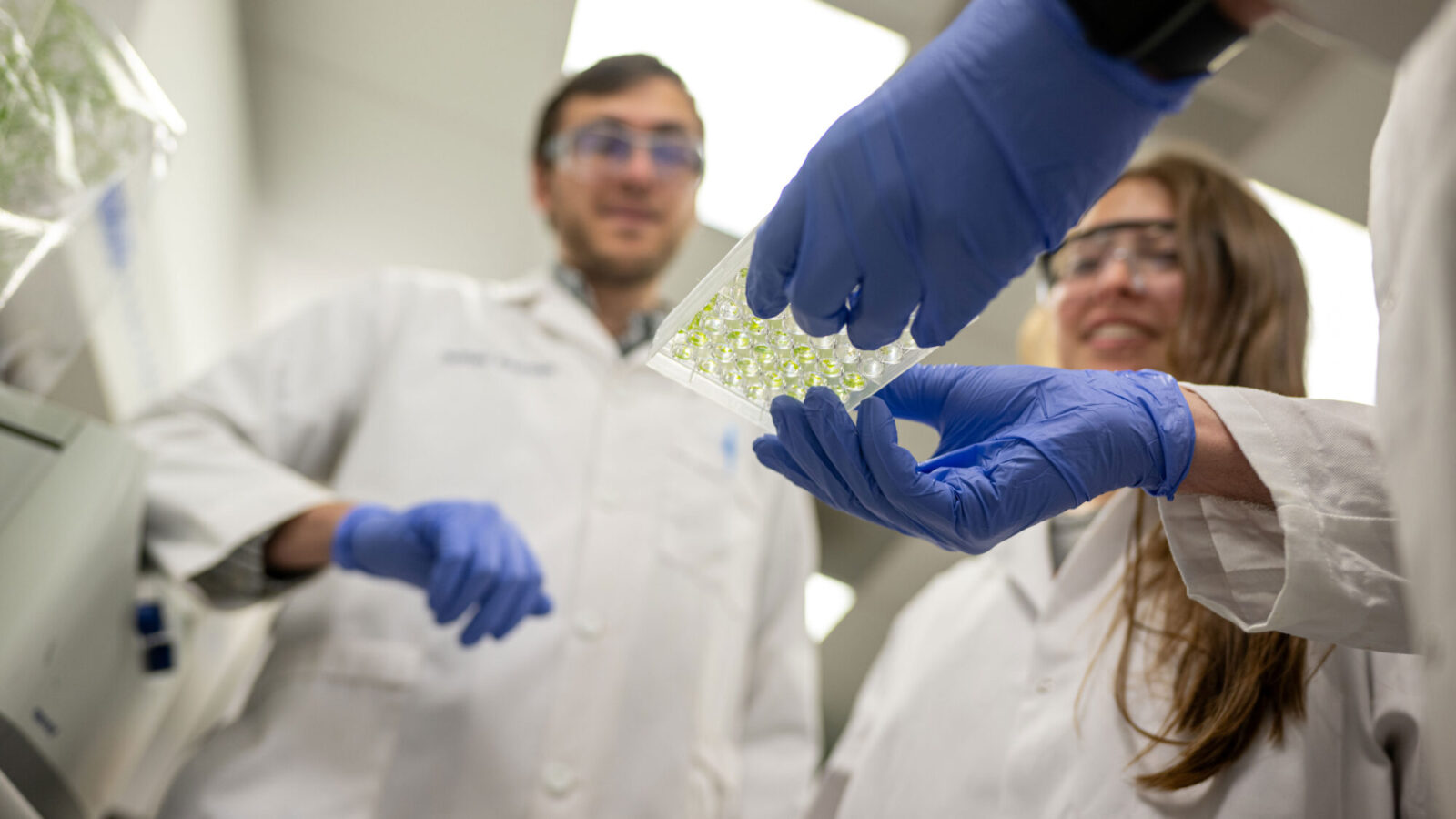
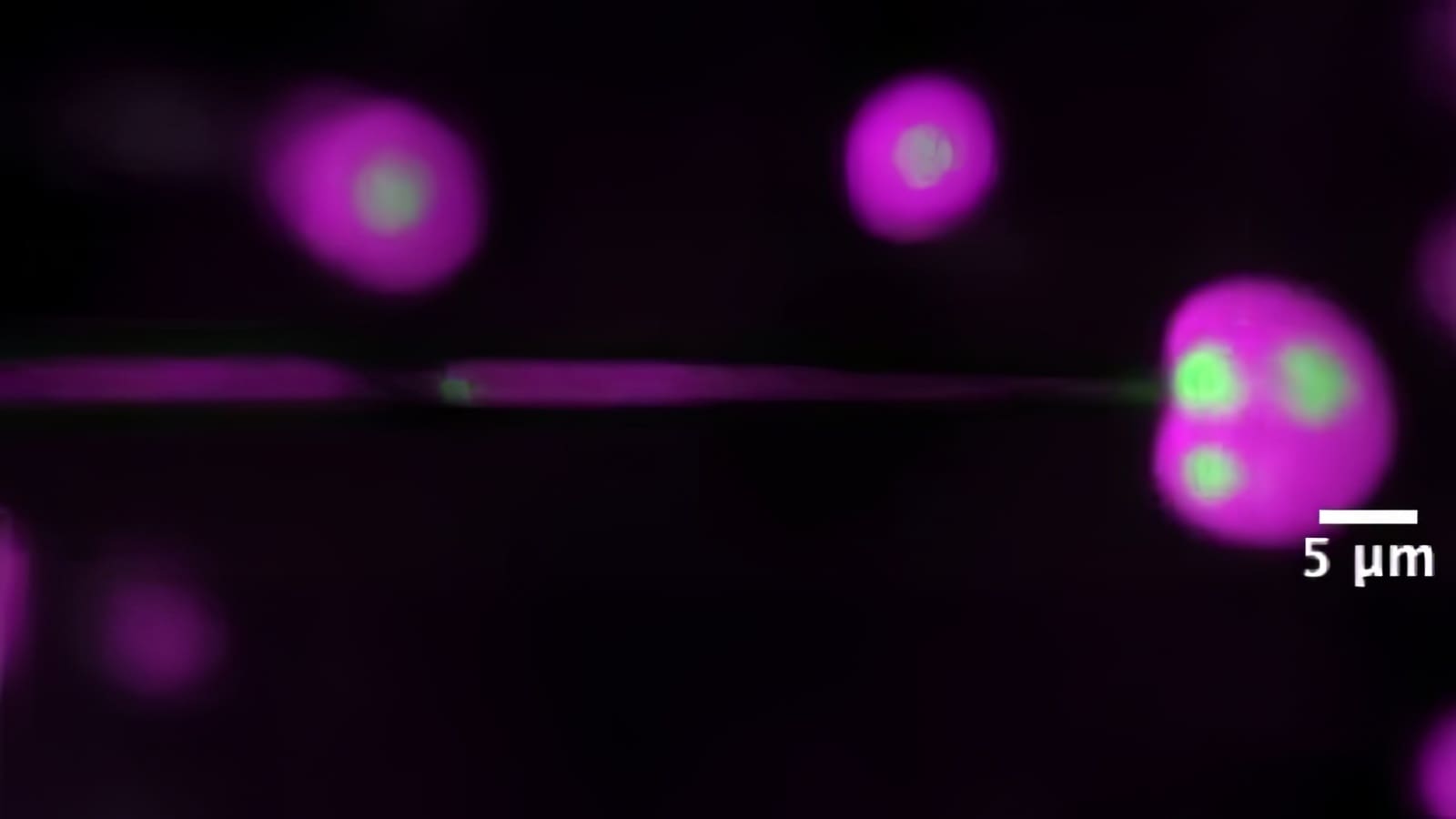
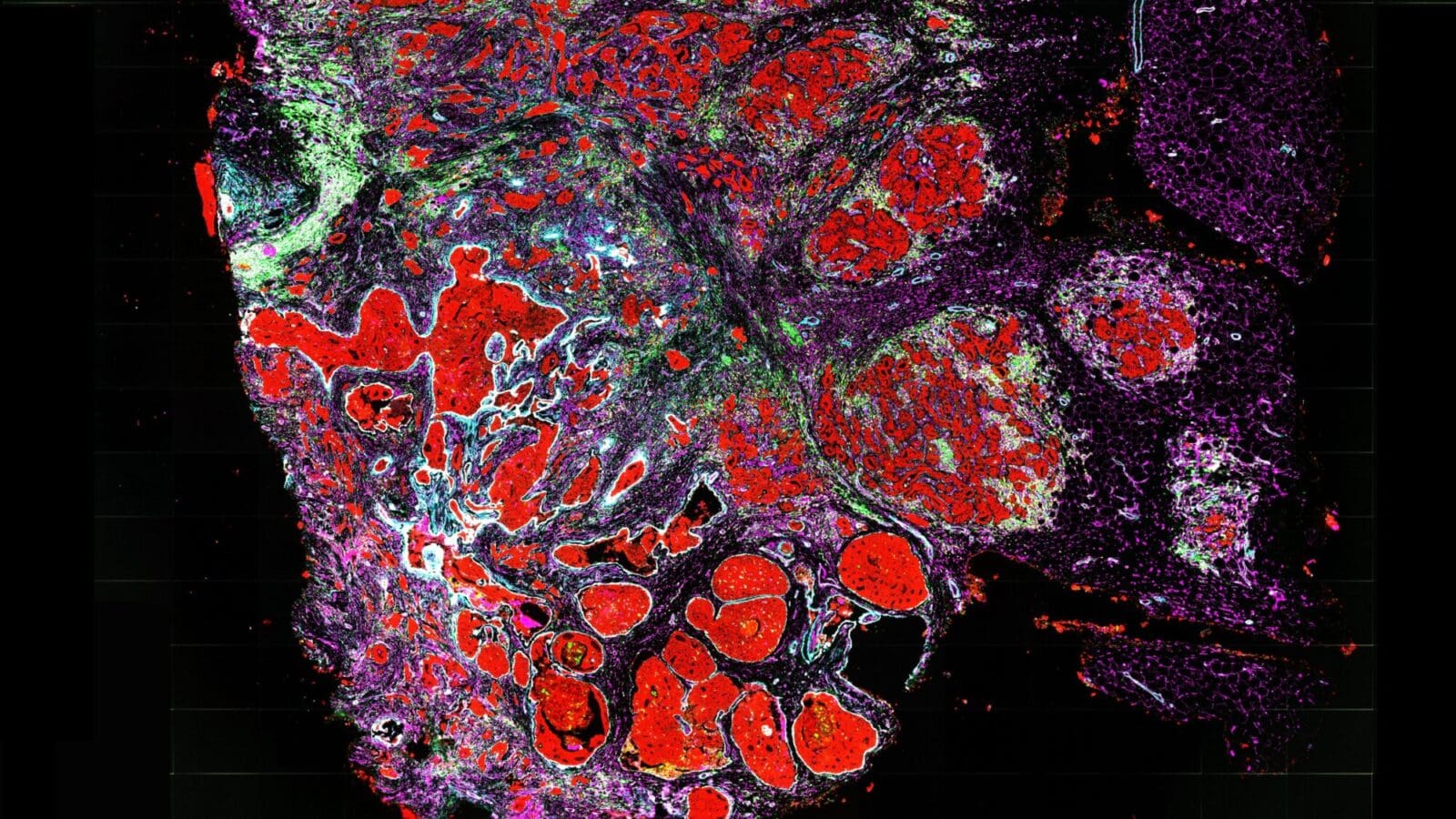
” title=”This computer-generated image shows a computationally designed drug, shown in yellow and
blue, binding to a cancer-related enzyme called EZH2, shown in red. Professor Christodoulos Floudas and
colleagues have created mathematical methods to predict, from billions of possibilities, what combinations
of amino acids would bind to naturally occurring proteins, thus leading to promising drug candidates.”
/>
Princeton researchers are applying Darwinian evolution principles to naturally occurring
antibacterial molecules to create novel antibiotics for the food and drug industries.
Bacteria secrete antimicrobial peptides — short chains of amino acids — for
defense against other species. James”>http://www.princeton.edu/cbe/people/faculty/link/”>James
Link, an assistant professor of chemical”>http://www.princeton.edu/cbe/”>chemical
and biological engineering, is pioneering research on a class of such peptides that are
lasso-shaped, which makes them resistant to the body’s defense mechanisms and hence good drug
candidates. “We’re understanding how these amazing structures are made by bacteria,”
Link said. “Thermodynamically they shouldn’t exist.”

style=”float: right; width: 200px; height: 275px; ” title=”Assistant Professor James Link and
graduate student Jessica (Si Jia) Pan are developing peptide drugs that could treat bacterial infections
that have become resistant to antibiotics. (Photo by Frank Wojciechowski)” />
Starting with one particular lasso peptide, Link and his graduate student Jessica (Si Jia) Pan have
created a dozen variants with more antibiotic potency. The researchers use a method called directed
evolution in which they create random mutations, test for desirable properties and repeat. They screened
20,000 variations of the peptide for the most promising molecules and found the most promising to be as
potent as the antibacterial peptides used in the food industry to protect perishables.
The researchers are now trying to beat bacteria at their own game. “We’re
trying to use directed evolution to find a peptide that can kill E. coli that are resistant to it,”
Link said. “In the same way that a bacteria evolves resistance, we can try to evolve peptides that
overcome that resistance.”
While Link’s approach to designing drugs is experimental, Christodoulos”>http://titan.princeton.edu/2010-10-11/People/floudas.html”>Christodoulos
Floudas, the Stephen C. Macaleer Professor of Chemical and Biological Engineering, has spent more
than a”>http://www.princeton.edu/engineering/news/archive/?id=4492″>a decade
developing computational methods for the same purpose.
Floudas and his graduate students have found a way to calculate whether a peptide, based on its
structure and amino-acid sequence, will bind to a specific protein. They have designed seven new peptides
that bind to a human enzyme linked to cancer progression. Experiments by a Boston company have shown that
the peptides inhibit the enzyme’s function. “There’s no currently available
inhibitor for this enzyme,” Floudas said.