Research into some bacteria that survive antibiotics, called persisters, has focused on the genes that allow some bacteria to tolerate antibiotic treatments. In their recent study, the Princeton researchers explored how the number of DNA copies in a cell affects whether a cell persists despite exposure to DNA-damaging antibiotics called fluoroquinolones. The study confirmed the researchers’ expectation that bacteria with backup chromosomal copies of DNA proved to be persisters at a much greater rate than cells with just one copy.

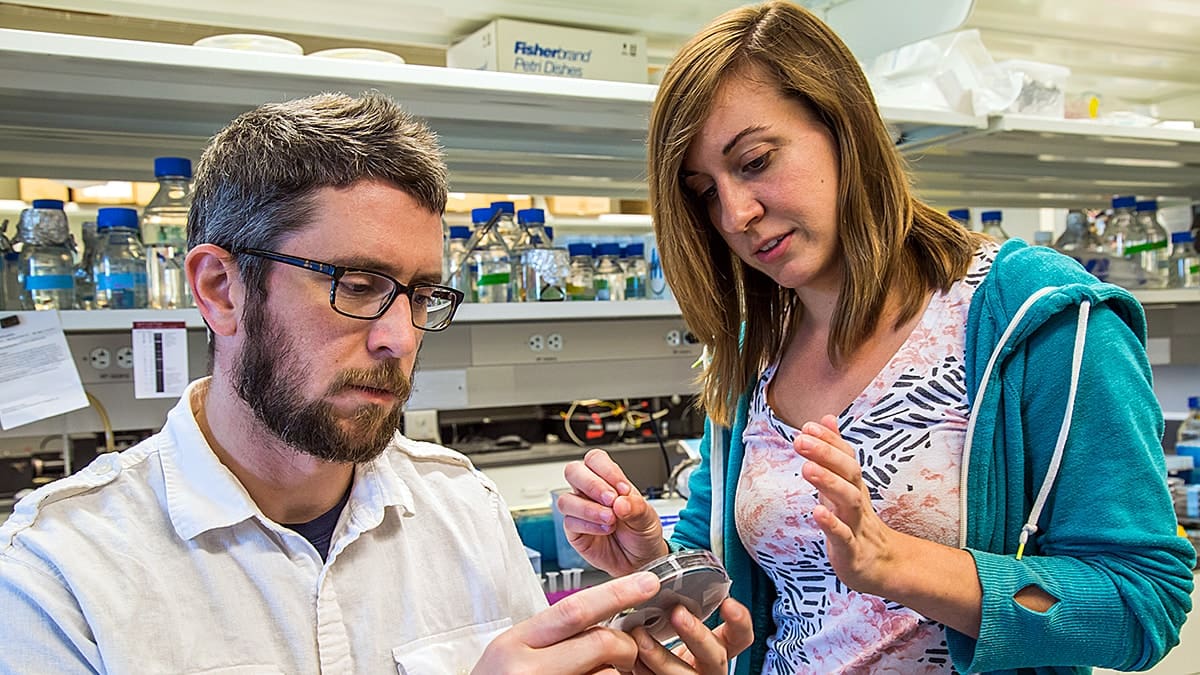
The study’s lead author Allison Murawski, an MD/Ph.D. student, started working on the project four years ago when she joined the lab of Mark Brynildsen, an associate professor of chemical and biological engineering at Princeton.
Aaron Nathans
Yet the results, which published March 11 in Current Biology, came with a significant twist. To a surprising degree, cells with just one DNA copy still managed to persist against fluoroquinolone treatment as well. The experiments revealed a second and separate pathway to persistence in cells with a lone DNA copy.
By bringing these two persistence pathways to light, the findings point the way to new strategies that could more effectively kill off the hardy microbes.
“Our study showed that there can be more than one type of persister to the same antibiotic in the same population,” said study lead author Allison Murawski, an MD/Ph.D. student who started working on the project four years ago when she joined the lab of Mark Brynildsen, an associate professor of chemical and biological engineering at Princeton.
“We had hypothesized that bacterial cells with only one chromosome wouldn’t be able to survive the antibiotic treatment,” said Murawski. “When we ran experiments confirming they do survive, that blew my mind.”
“Everything we had thought all along was just the opposite,” she added. “It’s those sorts of surprises and insights that make science really fun.”
“When Allison joined the lab, we thought this was going to be a quick project . . . but the bacteria had other ideas,” said Brynildsen, the study’s senior author. “As it turns out, its unfortunately not just a single pathway that the bacteria can take to fluoroquinolone persistence.”
Persisters have increasingly been recognized as a serious challenge to modern infection control. Unlike fully antibiotic resistant “superbugs,” persisters do not possess mutated genes; in fact, persisters are genetically identical to their normal, antibiotic-prone bacterial brethren. Persisters live up to their name, though, through the way those genes happen to be expressed at the time of an antibiotic onslaught. The persisters might be in a temporarily dormant state, for instance, or luckily have certain genes switched on or off that confer better protection from the antibiotic. Once the pharmaceutical threat has passed, persisters can start multiplying again and trigger a fresh round of infection.
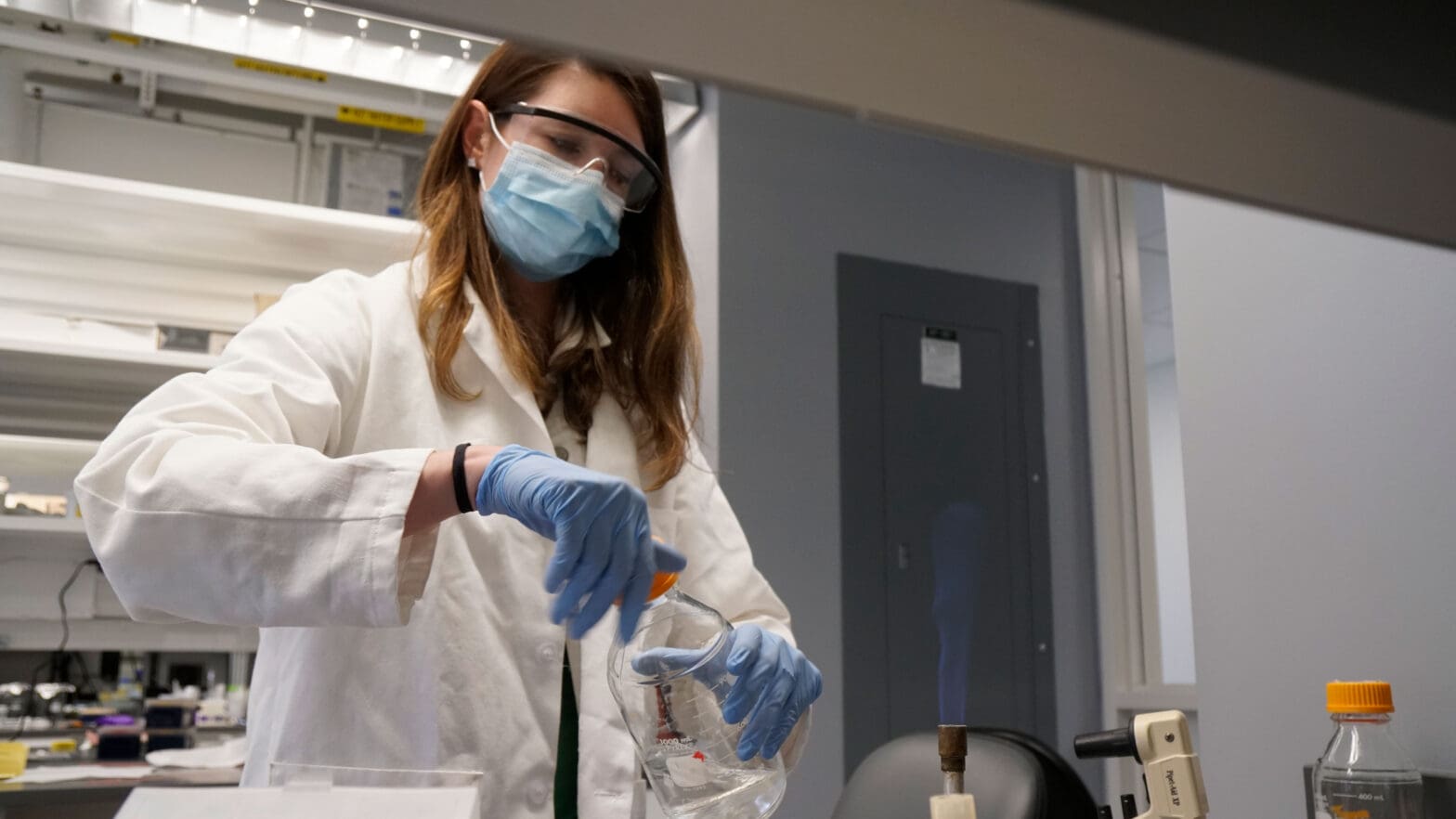
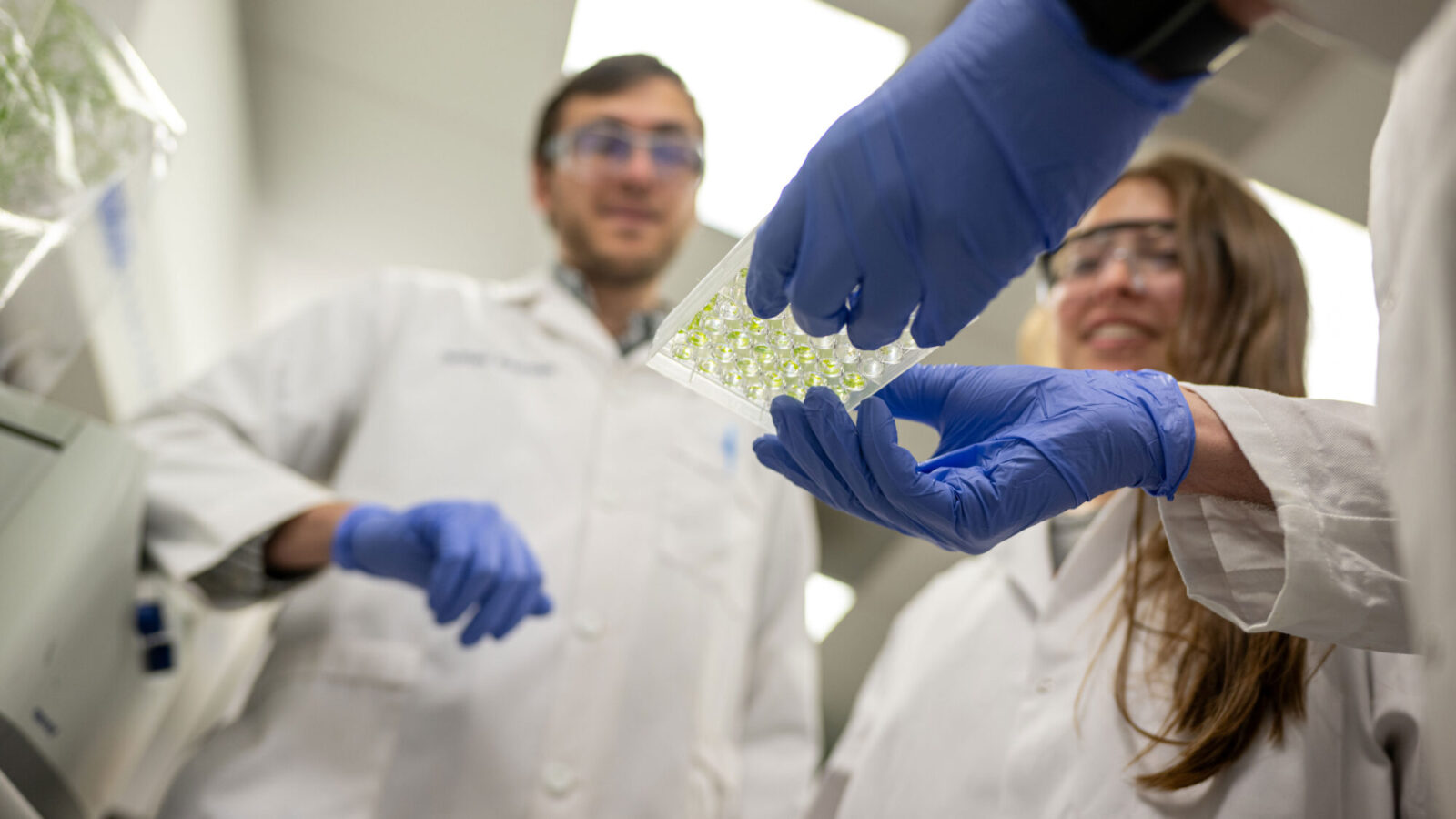
To probe the effect of the number of DNA copies, or ploidy, on persistence, Murawski and Brynildsen sorted cells of Escherichia coli bacteria into monoploid (single chromosome) and diploid (double chromosome) populations. When dosed with a fluoroquinolone called levofloxacin, most cells in each population died out. As hypothesized, however, the diploid bacterial cell populations harbored up to 40 times the number of persisters than the monoploid cell populations.
Having that extra DNA copy on hand enables the bacteria to perform homologous recombination—the term for when cells repair their broken DNA based on a template provided by an undamaged, additional chromosome. It’s like having a fully built Lego set to guide the rebuilding of a toppled set.
Digging deeper, the researchers created mutant E. coli cells that lacked one of two genes, dubbed RecA and RecB. These genes code for DNA-repair proteins that are required to perform homologous recombination. Sure enough, without the capacity to do homologous recombination, the mutant cells rarely endured as persisters, and it didn’t matter how many chromosomes they had.
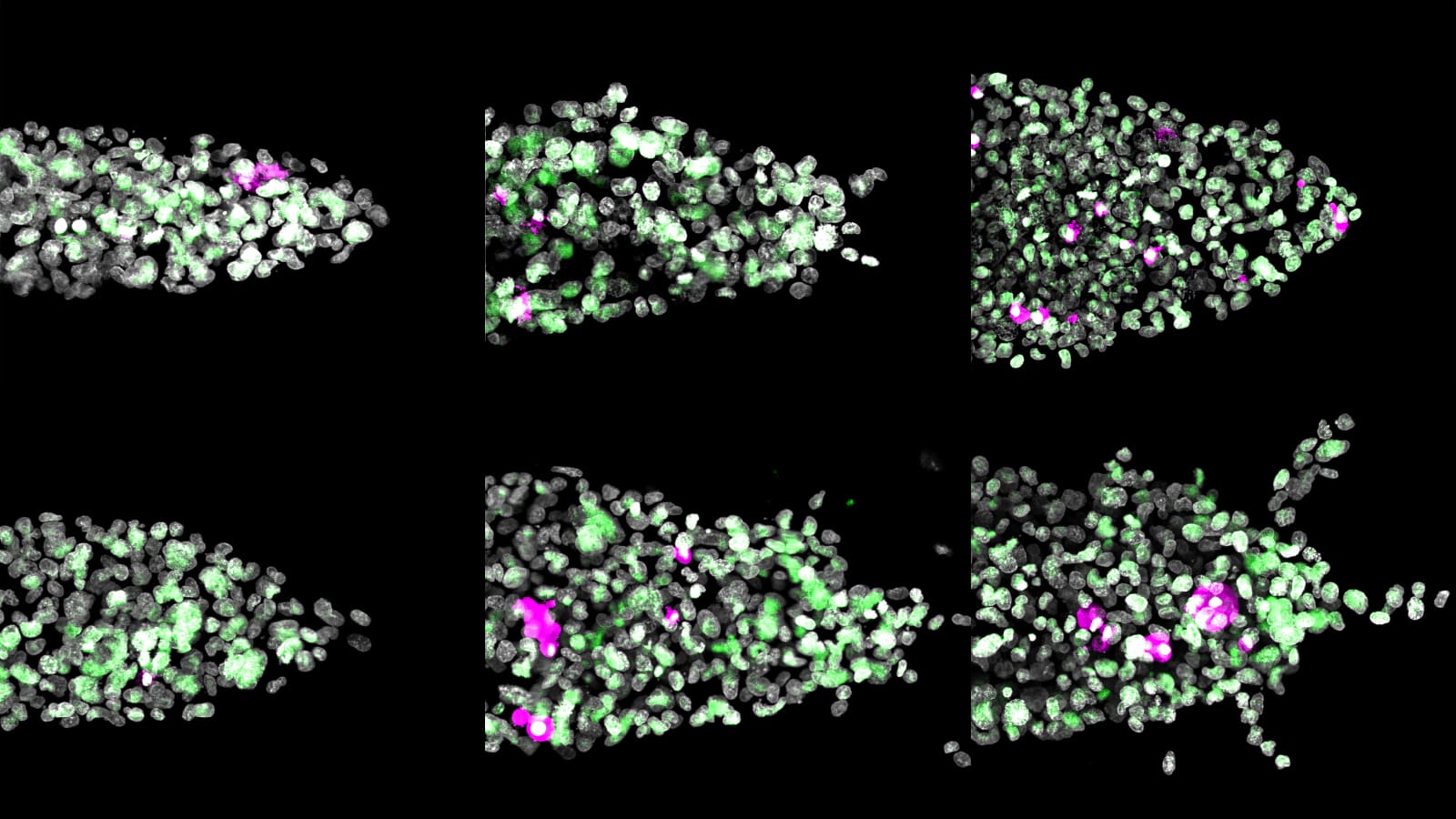
Puzzlingly however, persisters still cropped up in the monoploid wild-type cells, greater than 10-fold more often than the homologous recombination-deficient mutant cells. Without a backup DNA copy, it was not clear how the monoploid cells could repair their sundered DNA and live on. However, what was clear from live-cell imaging was that monoploid persisters recovered significantly different from treatment than their diploid counterparts, increasing in size more slowly and dividing later than cells with two chromosomes.
Given that single chromosome-bearing bacterial cells demonstrated a much lower propensity to persist, a novel kind of therapeutic agent suggested by the findings is one that fosters division of cells with multiple copies. For instance, if a drug promoted a diploid cell to divide, two monoploid cells would be generated, which in turn would be more vulnerable to fluoroquinolone treatment.
Meanwhile, RecA and RecB could prove to be suitable drug targets as well.
“With this line of research, we’re trying to identify ways to get rid of all of the persisters and successfully eradicate hard-to-treat infections,” said Brynildsen. “It looks as though we have our work cut out for ourselves.”
The work was supported by the National Institutes of Health.