Web of protein interactions governs cell behavior
 Mona Singh doesn’t use the maps in her office to get from point A to point B-she uses them to find meaning hidden in biological data, which may help advance the understanding of disease at the genetic level.
Mona Singh doesn’t use the maps in her office to get from point A to point B-she uses them to find meaning hidden in biological data, which may help advance the understanding of disease at the genetic level.
It’s a formidable task. The diagrams, called protein interaction maps, contain thousands of dots, each representing a different protein. A complicated network of lines connects the dots, diagramming how the proteins interact as they go about controlling everything the cell does. Though such maps have been generated for species ranging from yeast to humans, many of their insights remain obscured by the sheer volume of information they contain.
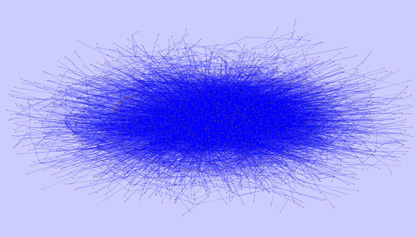

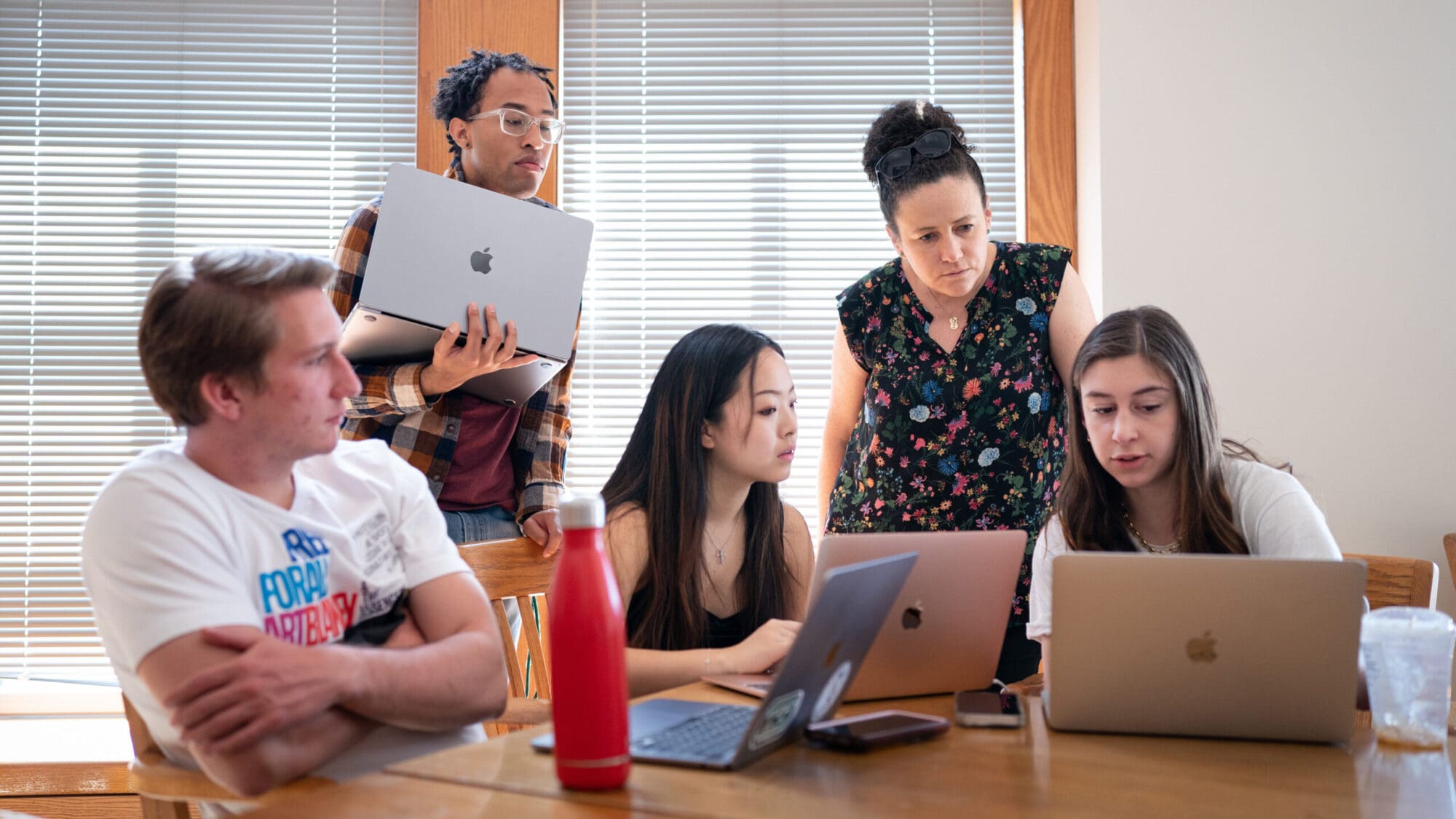
“We’re at an interesting point in terms of science,” said Singh, an associate professor of computer science and the Lewis-Sigler Institute for Integrative Genomics. “There is a lot of data being generated and you need to know fairly sophisticated computational methods to make sense of it.”
In partnership with Bernard Chazelle, professor of computer science, Singh designed a computer system that determines the function of unknown proteins using the known neighboring proteins in the map. For instance, if ten proteins known to be involved in cell pision all interact with an unknown protein, the mystery protein is likely to be involved in the division process as well.
Singh’s latest system detects patterns of interactions between proteins that occur more often than would be predicted by random chance. A pattern that is observed multiple times-e.g., the same types of proteins interacting-is likely to have functional significance as a cellular “building block,” and provides guidance for future biological investigation.
Singh also has been developing computational approaches to infer interaction maps and make high-confidence predictions of interactions that can be tested experimentally. This work is part of an ongoing collaboration with biologist Amy Keating at the Massachusetts Institute of Technology.