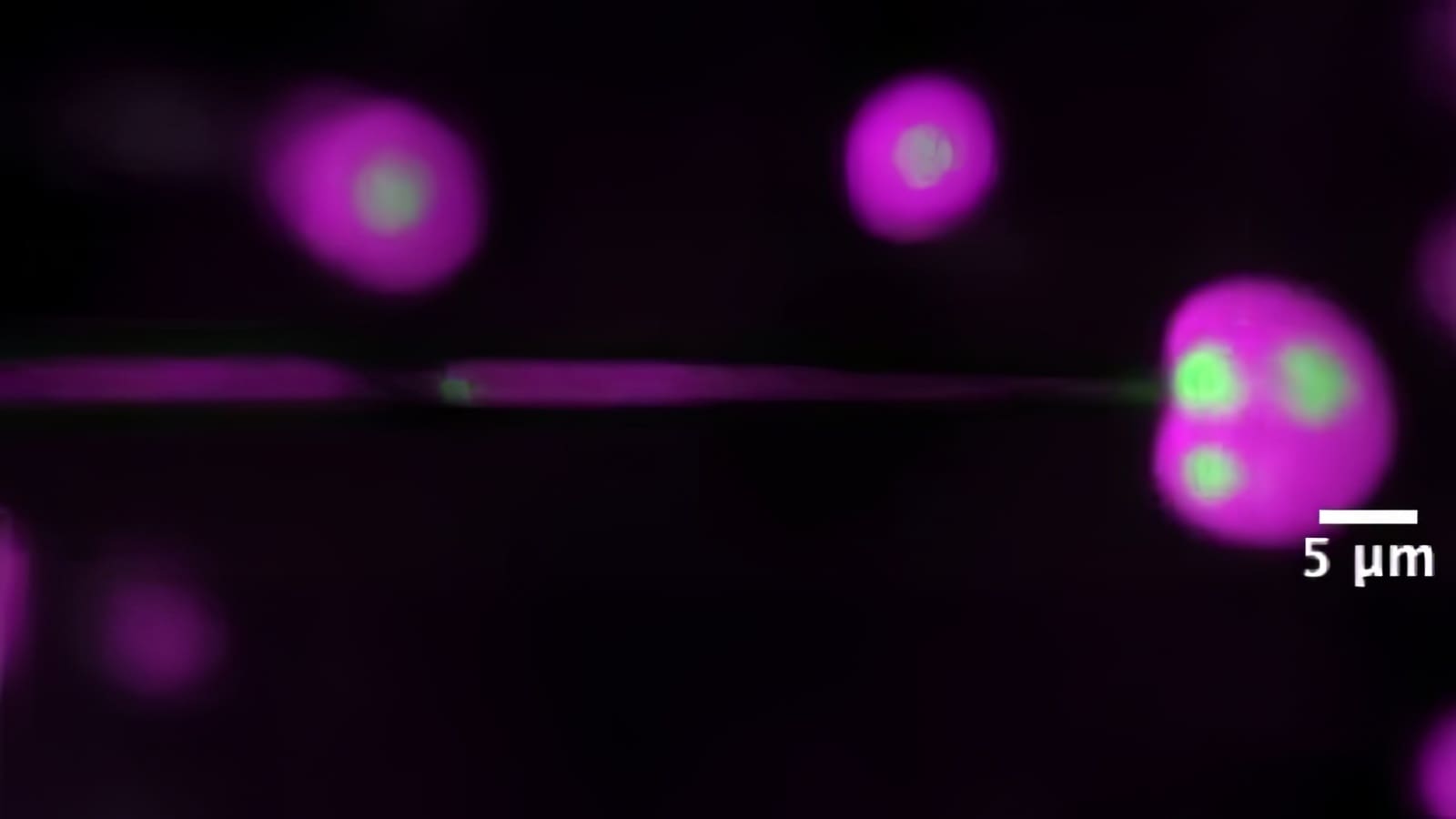
The discovery involves areas of DNA that do not directly code for the proteins that carry out cellular activities, but instead control how genes are switched on and off. Scientists have long understood that these noncoding regions play an important role in cancer, but describing that role has been formidably difficult. The team, led by researchers from Princeton University, said the discovery will open new lines of research about the development and spread of cancer.
“It’s really the beginning for understanding what’s happening in the noncoding space with cancer,” said Ben Raphael, a professor of computer science at Princeton and a senior author of the study, published Feb. 5 in Nature Communications. Other senior authors are J√ºri Reimand of the Ontario Institute for Cancer Research and Joshua Stuart of the University of California-Santa Cruz. The study’s lead author is Matthew Reyna, a former postdoctoral researcher at Princeton who is now at Emory University.
Pinpointing the genetic changes that lead to cancer is notoriously complex. With each tumor harboring thousands of individual mutations in its cells’ DNA, it can be difficult to distinguish “driver” mutations that enable cancer cells to grow and spread from “passenger” mutations that are simply along for the ride.

To find driver mutations, researchers typically analyze DNA sequences from large groups of patients, comparing sequences from cancer cells and healthy cells, and determine whether a mutation occurs significantly more often in cancer cells than would be expected by chance. But many driver mutations are relatively rare and are missed by this approach.
Raphael and his colleagues previously developed an algorithm to detect rarer driver mutations using knowledge of how groups of genes function together. Rather than analyzing one mutation at a time, the method assesses whether groups of genes that directly interact (networks) or participate in the same biological processes (pathways) are mutated more often in cancer cells than would be expected by chance. With an elaborate web of about 20,000 human genes, this is a formidable computational task.
In the current study, the researchers applied seven different pathway and network analysis methods to noncoding regions of whole genome sequences from more than 2,500 cancer patients – the largest collection to date of uniformly processed cancer genomes. Their paper is one of 23 studies published Feb. 5 in Nature journals as part of the Pan-Cancer Analysis of Whole Genomes project of the International Cancer Genome Consortium. The project involves more than 1,300 researchers who have used the data set to illuminate how noncoding portions of the human genome (which represent nearly 99% of its 3-billion-letter sequence) contribute to the development and progression of cancer.
Peter Campbell of the Wellcome Sanger Institute in the U.K., a member of the Pan-Cancer project steering committee, summed up the project’s contributions in a news release: “This work is helping to answer a long-standing medical difficulty: why two patients with what appear to be the same cancer can have very different outcomes to the same drug treatment. We show that the reasons for these different behaviors are written in the DNA. The genome of each patient’s cancer is unique, but there are a finite set of recurring patterns, so with large enough studies we can identify all these patterns to optimize diagnosis and treatment.”
The Princeton-led analysis focused on noncoding DNA regions that are part of genes but do not code for proteins. Instead, these sequences, known as promoters, enhancers and untranslated regions, act as signals to the cellular machinery that turns genes on and off. Using algorithms to search for mutations among genes within shared pathways and networks, the team uncovered 93 genes with potential driver mutations in their noncoding regions – only 19 of which were previously known cancer genes.
These genes are involved in processes that commonly go awry in cancer, such as cell proliferation and development. Other genes affect the status of chromatin, or the way DNA is packaged within the cell, which can markedly impact the expression of genes.
Perhaps the most interesting noncoding mutations found in the study were located near genes involved in RNA splicing, a key intermediate step that allows genes’ instructions to be translated into the proteins that carry out cellular activities. Relatively few RNA splicing genes contain cancer driver mutations in their coding sequences, but many are affected by noncoding mutations. The results bring up new research questions on the roles of RNA processing in cancer.
To test whether some of their predicted driver mutations affected actual gene activity patterns, Raphael and his colleagues turned to additional data from the Pan-Cancer project. From many of the patients, in addition to genome sequence data, the project had collected gene expression data – measuring which genes are active in a patient’s tumor. This allowed the researchers to demonstrate that the some of the newly predicted driver mutations actually changed how genes were turned on or off – albeit in a small number of samples containing mutations in a given gene.
“In a project like this, we have a tremendously rich data set, which allows us to perform all sorts of sophisticated algorithmic analyses, and make predictions,” said Raphael. “Then the big challenge is, what do you do with those predictions? Our work thus far has generated new hypotheses. The next step is experimental validation.”
One of the caveats to this type of analysis is that patterns of gene expression are highly specific to cell and tissue types, meaning that the same noncoding mutation could have different effects in lung cells than in breast cells, for example.
“Once you think about rare mutations, then 2,500 samples actually becomes extremely limiting,” said Raphael. “We expect to find only a handful of samples with mutations that occur in less than 1% of cancers.”
Still, Raphael says he is optimistic that other researchers will follow up on the team’s findings with new data and experiments.
Raphael’s work on this study was supported by the U.S. National Institutes of Health. Additional contributors included 22 co-authors from institutions in the United States, Canada and Europe, with contributions from the Drivers and Functional Interpretation Working Group of the Pan-Cancer Analysis of Whole Genomes project of the International Cancer Genome Consortium.