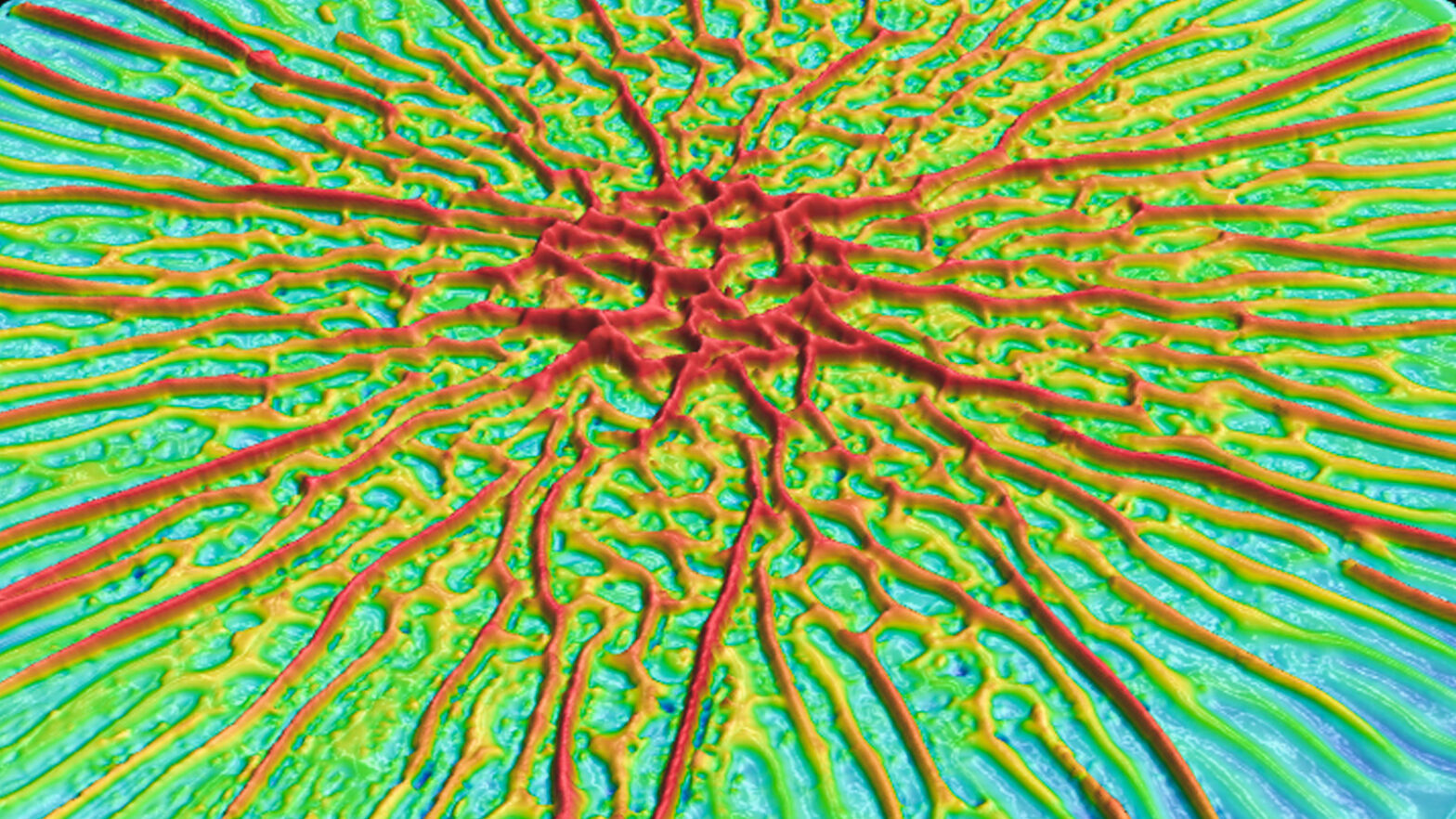
“At a certain point, patterns appear: stripes, zigzags and rings,” said co-author Ned Wingreen, the Howard A. Prior Professor in the Life Sciences and a professor of molecular biology and the Lewis-Sigler Institute for Integrative Genomics. “It’s been one of those things that has personally always puzzled me.”
The study, published Mar. 19 in the Proceedings of the National Academy of Sciences, helps illuminate how bacteria grow in diverse environments, and could also aid investigations into the physical forces that govern growth and morphing of human tissues.
“This adds to a body of work coming from a mechanical perspective that says what we’re seeing is the playing out of physical laws,” said Wingreen. “It will help us understand to what extent some of these patterns influence the biofilm properties that are important biologically and medically.”
The work is the latest to stem from a collaboration among four faculty members investigating biofilms from multiple perspectives. Along with Wingreen, the team includes Andrej Ko≈°mrlj, an assistant professor of mechanical and aerospace engineering; Howard Stone, the Donald R. Dixon ’69 and Elizabeth W. Dixon Professor of Mechanical and Aerospace Engineering; and Bonnie Bassler, the Squibb Professor in Molecular Biology. They previously worked together to develop a method for peeling biofilms off surfaces.
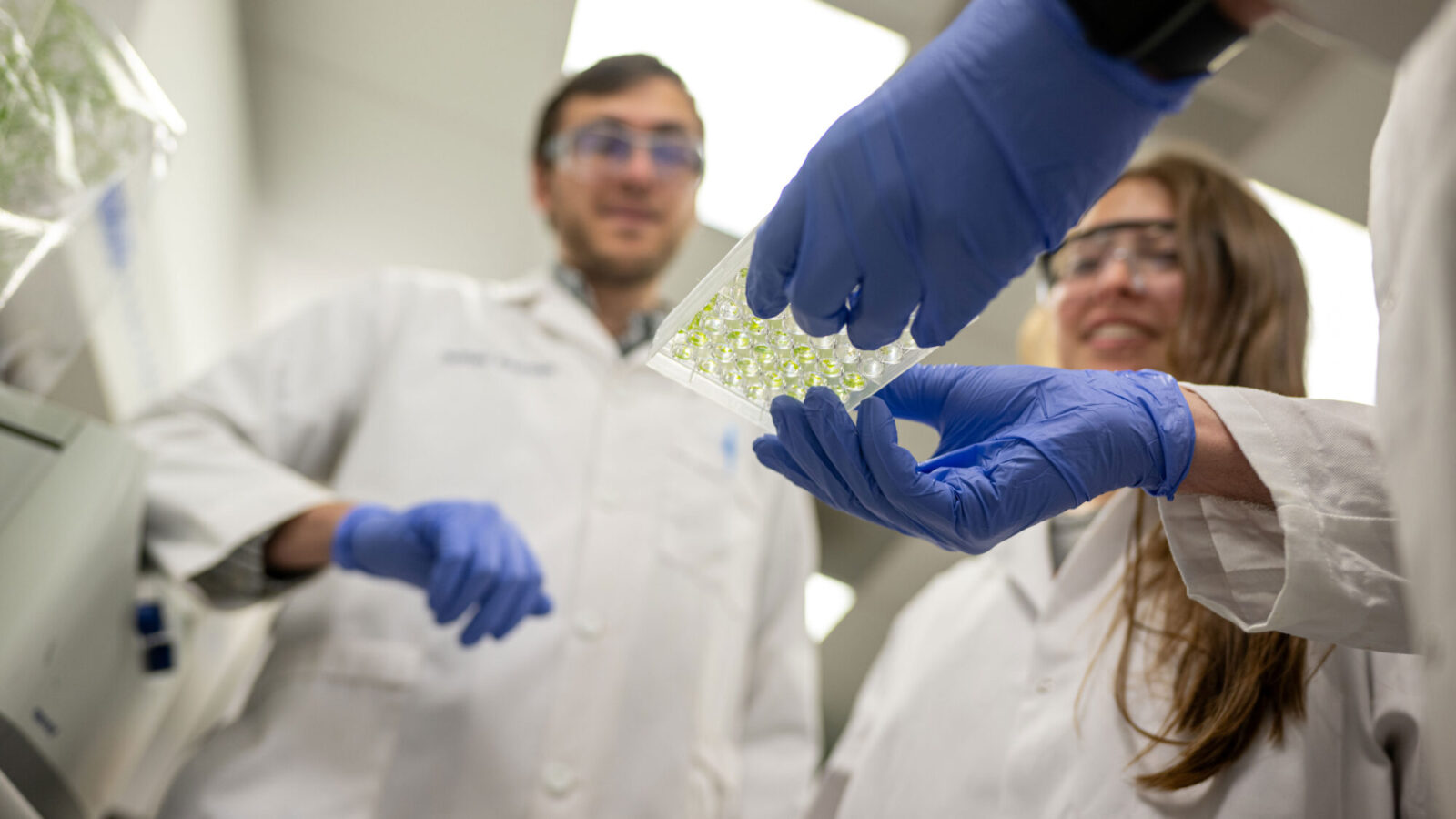
In this study, the researchers analyzed biofilm formation of the bacterium Vibrio cholerae, which is widespread in watery environments and can cause the disease cholera when ingested by humans. On a soft substrate, the biofilms initially grow as a flat layer but later become wrinkled, with radial and zigzag patterns of peaks and valleys. The study showed that these patterns depend on the relative softness or stiffness of the biofilm’s substrate, which the researchers manipulated by varying the concentration of agar, a seaweed-derived substance commonly used to fill Petri dishes.
On a stiffer substrate, wrinkles first appear at a biofilm’s center and propagate outward, while on a softer substrate wrinkles begin to form at the edges and propagate toward the center. In both cases, biofilms end up with zigzag wrinkle patterns at their centers and a more ordered pattern of radial stripes at their outer edges.
Time-lapse video showing the growth of a V. cholerae biofilm on 0.7% agar. On this stiff substrate, wrinkles first appear in the center and propagate outward. Video from eLife 2019;8:e43920 DOI: 10.7554/eLife.43920
This process is driven by a nexus of influences, including each bacterium’s uptake of nutrients from the substrate, which leads to uneven growth as nutrients are depleted in the biofilm’s center. The bacteria produce new extracellular matrix components as they grow and they also produce molecules that make the bacteria stick to each other and to the substrate.
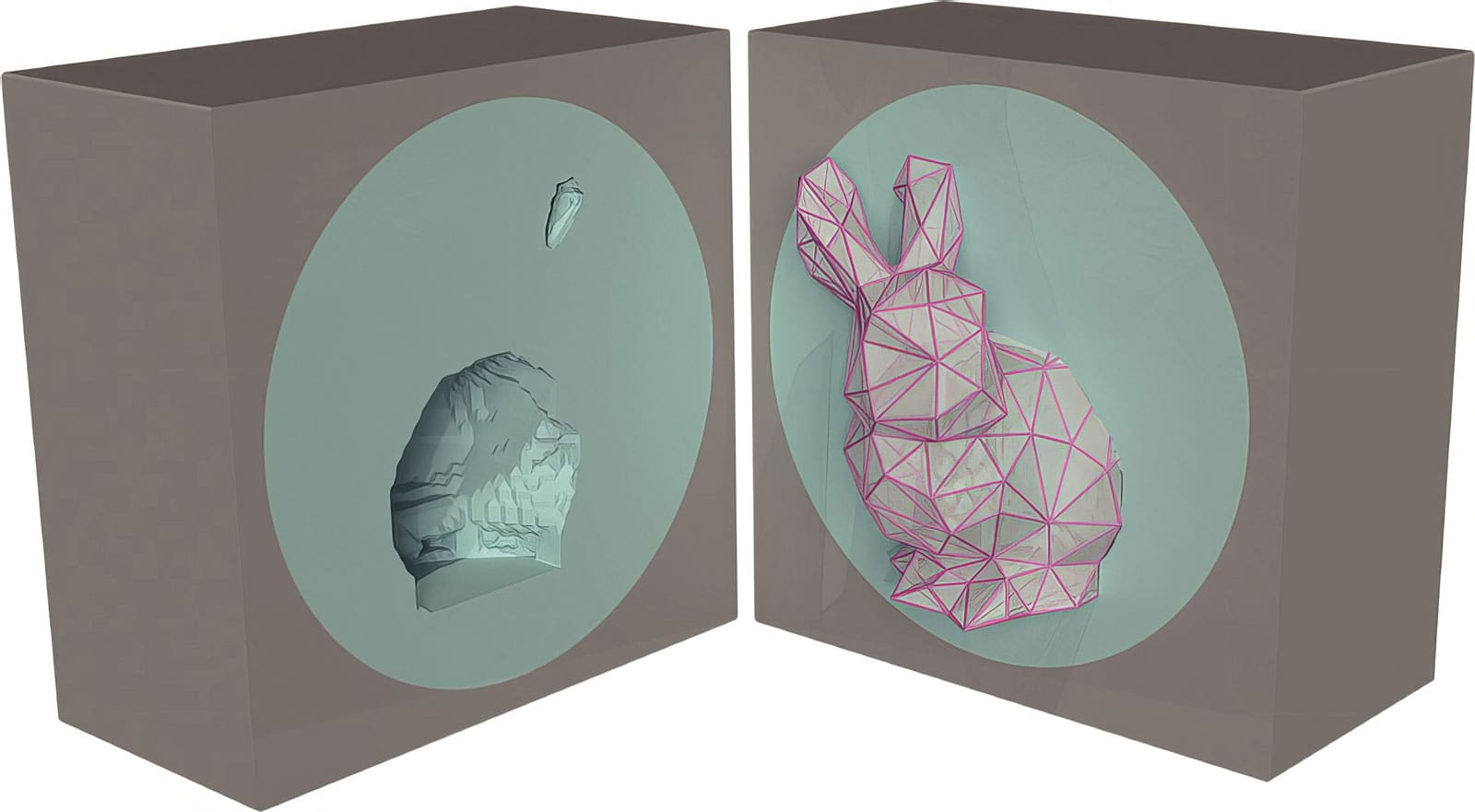
“This is a very complicated process involving growth and mechanics,” said lead author Chenyi Fei, a graduate student in the Lewis-Sigler Institute. “To understand it, we built what we call a chemo-mechanical model. We take into account the nutrients and the nonuniform growth of the biofilm, and how those features translate into the mechanical forces or stresses that accumulate.”
The model allowed the researchers to predict which regions of the biofilm would experience maximum stress at particular points in time, and thus to predict where wrinkles would form. The predictions corresponded well with experimental measurements of three-dimensional shapes of actual biofilms, said Fei, whose graduate work is co-advised by Wingreen and Bassler.
“Chenyi cleverly integrated a lot of different mechanisms that are normally studied separately,” said Ko≈°mrlj.
Time-lapse video showing the growth of a V. cholerae biofilm on 0.4% agar. On this soft substrate, wrinkles begin to form at the biofilm’s edges and propagate toward its center. Video from eLife 2019;8:e43920 DOI: 10.7554/eLife.43920
A key aspect of the model is its adoption of classic engineering analyses of mechanical instabilities, which have previously been applied to such problems as the buckling of railroad tracks in extreme temperatures. A similar type of instability causes soft materials like biofilms to form wrinkles.
“In the previous century, mechanical instabilities were being studied with a focus on trying to prevent failure mechanisms in structures,” said co-author Sheng Mao, a former postdoctoral researcher at Princeton who is now an assistant professor at Peking University. “But in a new wave of studies, we are trying to exploit these mechanical instabilities to make tunable structures for various purposes,” such as soft materials being developed to treat injuries and diseases through tissue engineering.
In addition to advancing knowledge of how growing cells interact with soft substrates, the researchers plan to build on their findings to further investigate bacterial life cycles, including the dispersal stage in which some bacterial cells detach from a biofilm, exit the structure, and colonize new areas.
Bacteria such as V. cholerae “are opportunists,” said Wingreen. “They come in, grab some territory, expand, eat everything they can, and then leave. But this last aspect, dispersal, is understudied, and the physics and mechanics of dispersal are going to offer some interesting challenges.”
In addition to Wingreen, Ko≈°mrlj, Stone, Bassler, Fei and Mao, other study co-authors were Jing Yan, a former postdoctoral researcher who is now an assistant professor at Yale University; and Ricard Alert, a postdoctoral fellow in Princeton’s Center for Theoretical Science.
This work was supported in part by the Howard Hughes Medical Institute, the U.S. National Institutes of Health and the National Science Foundation, including an NSF award to the Princeton Center for Complex Materials.